Research article
Synthesis and computational investigation of molecularly imprinted nanospheres for selective recognition of alpha-tocopherol succinate
Theeraphon Piacham1[*], Chanin Nantasenamat1,2, Chartchalerm Isarankura-Na-Ayudhya1, Virapong Prachayasittikul1
1Department of Clinical Microbiology and Applied Technology, Faculty of Medical Technology, Mahidol University, Bangkok 10700, Thailand2Center of Data Mining and Biomedical Informatics, Faculty of Medical Technology, Mahidol University, Bangkok 10700, Thailand
EXCLI J 2013;12:Doc701
Abstract
Molecularly imprinted polymers (MIPs) are macromolecular matrices that can mimic the functional properties of antibodies, receptors and enzymes while possessing higher durability. As such, these polymers are interesting materials for applications in biomimetic sensor, drug synthesis, drug delivery and separation. In this study, we prepared MIPs and molecularly imprinted nanospheres (MINs) as receptors with specific recognition properties toward tocopherol succinate (TPS) in comparison to tocopherol (TP) and tocopherol nicotinate (TPN). MIPs were synthesized using methacrylic acid (MAA) as functional monomer, ethylene glycol dimethacrylate (EGDMA) as crosslinking agent and dichloromethane or acetronitrile as porogenic solvent under thermal-induced polymerization condition. Results indicated that imprinted polymers of TPS-MIP, TP-MIP and TPN-MIP all bound specifically to their template molecules at 2 folds greater than the non-imprinted polymers. The calculated binding capacity of all MIP was approximately 2 mg per gram of polymer when using the optimal rebinding solvent EtOH:H2O (3:2, v/v). Furthermore, the MINs toward TPS and TP were prepared by precipitation polymerization that yielded particles that are 200-400 nm in size. The binding capacities of MINs to their templates were greater than that of the non-imprinted nanospheres when using the optimal rebinding solvent EtOH:H2O (4:1, v/v). Computer simulation was performed to provide mechanistic insights on the binding modalities of template-monomer complexes. In conclusion, we had successful prepared MIPs and MINs for binding specifically to TP and TPS. Such MIPs and MINs have great potential for industrial and medical applications, particularly for the selective separation of TP and TPS.
Keywords: molecular imprinting, molecularly imprinted polymer, anti-cancer, tocopherol succinate, computational chemistry
Introduction
Significant changes to the environment and climate as a result of global warming had increased the exposure to toxic substances that may culminate in the development of pathogenic diseases (Dapul-Hidalgo and Bielory, 2012[15]; Hunter, 2003[24]; Thomas et al., 2012[68]). Among these, cancer has been found to increase incidentally owing to increases of UV exposure and toxicant-induced gene mutation. The development of therapeutic agent toward cancer has predominantly focused on addressing issues pertaining to its toxicity, drug delivery properties and multidrug resistance (Abraham et al., 2000[1]; Dubikovskaya et al., 2008[17]). Furthermore, intense efforts have been invested in improving therapeutic approaches as to increase patient survival (Bechet et al., 2008[4]; Campbell et al., 2009[7]).
Tocopherol succinate (TPS), a vitamin E analogue, is a promising and attractive compound with known anti-cancer activity toward several types of human cancer cell lines. Particularly, TPS can selectively induce apoptosis in malignant cells (Constantinou et al., 2008[13]; Neuzil, 2003[43]; Shanker et al., 2007[61]; Zhao et al., 2009[79]) while being non-toxic to normal cells and tissues. Structure-function relationship study of the terminal dicarboxylic moiety of tocopherol (TP) analogues have been previously investigated (Kogure et al., 2004[26]) and it was concluded that the apoptogenic activity depended on the length and charge of the ester moiety. Birringer et al. (2003[6]) provided further insights into the structure-function relationship of vitamin E by dividing the structure into three distinct domains. The pharmacokinetic property of TPS is similar to that of TP in which after infusion it is circulated in the blood stream by docking to lipoproteins where it subsequently targets the micro-capillary of tumor cells. In regards to its physicochemical properties, the hydrophobic nature of the molecule is responsible for the propensity of TP to bind lipoprotein and travel through the peripheral tissues followed by its sequential transfer to tumor cells. As compare to the normal tissue that exerts neutral state membrane, malignant cells possess acidic membranes in the protonated state. The inherent physicochemical property of TPS enables it to counteract this by being freely diffusible into malignant cells owing to its weak acidic nature that comprises of charged and deprotonated moieties. TPS undergoes hydrolysis and is converted to TP by nonspecific esterases from hepatocytes (Neuzil and Massa, 2005[44]; Wu and Croft, 2007[74]).
Molecular imprinting is a technique that affords the production of synthetic receptors or so-called plastic antibodies. Such molecularly imprinted polymers (MIPs) are recognition matrices that have the ability to recognize and bind specifically to compounds of interest. MIPs are known to possess higher durability than biological receptors as it is known to possess excellent thermostability, reusable and is easy to store (Bagheri et al., 2012[3]). As such, MIP has been successfully utilized for various applications such as substitutes for biological antibodies and receptors (Ye and Mosbach, 2008[76]), separation matrices for chromatography (Wei et al., 2005[71]) and solid phase extraction (Pichon and Haupt, 2006[52]), analytical sensors (Piacham et al., 2005[50]; Ton et al., 2012[69]), immuno assays (Moreno-Bondi et al., 2012[33]), drug delivery (Cunliffe et al., 2005[14]; Puoci et al., 2011[59]), enzyme inhibitor synthesis (Yu et al., 2002[77]; Zhang et al., 2006[78]) and enzyme mimetics (Piacham et al., 2003[48], 2006[49]). In recent years, molecular imprinting have been employed in the synthesis of polymers affording robust recognition properties toward several compounds of interests and several recent reviews have been published describing their potential utilizations for selective separation of compounds (Chen and Li, 2012[9]; Cheong et al., 2013[10][11]; Hu et al., 2013[23]; Murray and Örmeci, 2012[34]; Sharma et al., 2012[62]; Vasapollo et al., 2011[70]). Of particular note, is its utilization as specific drug delivery agents such as for metal-based anti-inflammatory drug (Sumi et al., 2008[66]), glycyrrhizic acid (Cirillo et al., 2010[12]), hyaluronic acid (Ali and Byrne, 2009[2]) and 5-fluorouracil (Puoci et al., 2007[60]) among others.
Molecularly imprinted polymers are prepared in essentially three major steps:
(i) Formation of template-monomer complexes, cross-linking and polymerization (Komiyama et al., 2003[27]). The first step involving the self-association of template-monomer complexes in which functional monomers affording the strongest binding are typically deemed as the optimal one to use in molecular imprinting experiments. Several approaches exist for calculating the interaction energy of template-monomer complexes:
(ii) Computational chemistry approach (i.e. involving the computation of the molecular energy of template, monomer and their complexes followed by calculating their energetic difference) (Piletsky et al., 2001[53]; Subrahmanyam et al., 2001[63]),
(iii) Data mining approach (i.e. involving the use of multivariate analysis methods such as neural network for correlating molecular descriptors with their experimental imprinting factor) (Nantasenamat et al., 2007[37], 2005[41]), molecular dynamics approach (i.e. involving the construction of atomistic model of molecularly imprinted polymer in which the cross-linked polymer contained template cavities to mimic experimental settings) (Dong et al., 2009[16]; Herdes and Sarkisov, 2009[22]; Lv et al., 2008[31]). Previously, we had successfully employed the computational chemistry approach for calculating the interaction energy of sulfonamides (Isarankura-Na-Ayudhya et al., 2008[25]) and tocopherols (Piacham et al., 2009[51]). The data mining approach was first introduced by us as a facile method that allows simultaneous modeling of molecular and solvent descriptors by means of quantitative structure-property relationship (QSPR) to make predictions of the imprinting factor values for template-monomer complexes of interests (Nantasenamat et al., 2007[37], 2005[41]). A more extensive account on computational approaches for the rational design of molecularly imprinted polymers has previously been reviewed (Levi et al., 2011[29]; Nicholls et al., 2009[45], 2011[46]). Further information on QSPR is provided in our previous review articles (Nantasenamat et al., 2009[35], 2010[38]) and the utilization of such modeling approach had successfully been demonstrated on a wide range of biological activities (Mandi et al., 2012[32]; Pingaew et al., 2012[54]; Thippakorn et al., 2009[67]; Worachartcheewan et al., 2011[72], 2009[73]) and chemical properties (Nantasenamat et al., 2007[37][39], 2008[36], 2005[41], 2013[42]).
A wide range of MIPs with specific binding properties towards TP and its derivatives has been synthesized by various polymerization techniques. Faizal and Kikuchi, (2010[19]) utilized molecular imprinting membranes synthesized via phase inversion technique bearing calix[4]resorcarenes moieties that engages in multiple non-covalent interactions with TP. The imprinted membrane was reported to bind TP over 2-folds higher than the non-imprinted membranes in methanol/water (2:1, v/v). Furthermore, Faizal et al. (2008[18]) had also employed the phase inversion imprinting technique for preparing copolymers of acrylonitrile with α-tocopherol methacrylate (TPM) monomer by means of the covalent imprinting method. Moreover, Faizal and Kobayashi (2008[20]) had also synthesized a hybrid molecular imprinting polymer for TP using pre-polymerization powders of TPM cross-linked by divinylbenzene onto a scaffold matrix of polysulfone, cellulose acetate, and nylon. Puoci et al. (2007[58]) demonstrated the use of α-TP imprinted polymer as solid phase extraction sorbents of α-TP from bay leaves extract. In our previous investigation we had successfully prepared bulk monoliths and nanospheres for selective recognition of TP and TP acetate (TPA). The study also provided mechanistic insights into the binding modes of TP-MAA and TPA-MAA complexes (Piacham et al., 2009[51]). Liu et al. (2012[30]) prepared MIP-based nano-sensing material towards TP by anchoring the MIP layer onto 1-vinyl-3-octylimidazolium ionic liquid-modified CdSe/ZnS quantum dots. Such smart material exhibited florescence quenching signal upon TP binding without the need for inducers and derivatization. TP-MIP as drug delivery devices has been investigated for their recognition characteristics in organic and aqueous media including their release property in gastrointestinal simulated fluids (Puoci et al., 2008[57]).
Molecularly imprinted polymers with binding specificity towards the apoptogenic TPS have not yet been reported. Therefore, this study aims to synthesize bulk monoliths and nanospheres via precipitation polymerization for further application as TPS delivery matrices. Computer simulation was also employed to provide pertinent insights on the binding modalities of the investigated template-monomer complexes in order to understand the origins of the observed binding performance.
Materials and Methods
Reagents
Tocopherol (TP), tocopherol nicotinate (TPN), tocopherol succinate (TPS), methacrylic acid (MAA), ethyleneglycol dimethylacrylate (EDMA) and azobis-isobutyronitrile (AIBN) were purchased from Sigma-Aldrich. All solvents were of analytical or HPLC grade.
Preparation of molecularly imprinted polymer
Molecularly imprinted polymers were synthesized using TP, TPN and TPS as template molecules (Figure 1(Fig. 1)). To 10 ml of DCM, 0.5 mmol of template, 8 mmol of MAA as functional monomer, 50 mmol of EDMA as cross-linking monomer and 202 mg of AIBN were added. The pre-polymerization mixture was purged with argon for 10 min and screw-capped in a 20 ml borosilicate tube. Thermal-induced polymerization was performed in a pre-heated water bath at 60 °C for 18 h. The obtained monolithic polymer was ground by a mechanical mortar to produce particles of varying sizes. Sedimentation in acetone was performed to separate and collect particles with diameters around 10-25 μm. Templates were eluted from the polymer by using acetic acid: methanol (15 %, v/v). Spectrophotometric examinations were performed in order to determine the amount of remaining templates. Non-imprinted polymers (NIPs) were prepared in the same manner with the omission of templates.
Preparation of molecularly imprinted polymers nanospheres
Molecularly imprinted polymers nanospheres (MINs) toward TP and TPS were prepared according to Ye et al. (1999[75]). Briefly, 80 ml of pre-polymerization mixture in acetonitrile contained 0.5 mmol of template, 8 mmol of MAA, 50 mmol EDMA and AIBN. The pre-polymerization mixture was purged with argon for 15 min prior to subjecting it to thermal-induced polymerization at 60 °C for 24 h. Templates were then eluted from the obtained nanospheres using acetic acid:methanol (15 %, v/v) via Soxhlet extraction. The non-imprinted polymers nanospheres (NINs) were prepared in the absence of template molecules.
Scanning electron microscopy
Particle sizes of imprinted and non-imprinted nanospheres were determined from scanning electron microscope (HITACHI S-3400). Briefly, the nanospheres were mounted on metallic studs via double-sided conductive tape and subsequently applying gold ion coating using sputter coater (Bal-tec SCD 050) for 90 s under vacuum at current intensity of 60 mA and scanning accelerating voltage of 15 kV.
Binding analysis
Binding analysis was performed by incubating a fixed amount of template (0.1 mg/mL) against various amounts of polymers in a 1 ml microfuge tube on a rocking table at room temperature for 12 h. Supernatants were taken after centrifugation at 12,000 rpm for 5 min prior to determining the amount of template bound via spectrophotometry.
Computer simulation
Chemical structures of template molecules (i.e. TP, TPN and TPS) and functional monomer MAA were drawn into Marvin Sketch (ChemAxon Ltd.) and were subsequently converted to the Gaussian input file format using Babel (OpenEye Scientific Software). Full geometry optimizations with no symmetry constrains were then performed initially at the semi-empirical AM1 level in order to afford good starting structures for subsequent refinement at the density functional theory level using Becke's three-parameter Lee-Yang-Parr (B3LYP) (Becke, 1993[5]; Lee et al., 1988[28]) and the 6-31G(d) basis set. Single point energy calculation was then performed on these optimized structures using the B3LYP functional and 6-311++G(d,p) basis set. All computational chemistry calculations were performed using Gaussian 09 (Frisch et al., 2009[21]).
Template-monomer complexes were then constructed by using the optimized structures of template and monomer. The functional monomer MAA was placed near each of the oxygen and nitrogen atoms of the template molecule to give rise to several possible template-monomer complexes. These structures were then subjected to full geometry optimizations. Finally, the total energy of the template molecules, functional monomers and complexes were parsed from the Gaussian output files using in-house developed scripts. The interaction energy was then computed using the following equation:
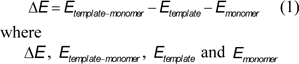
represents the interaction energy, energy of template-monomer complexes, energy of template molecule and energy of functional monomer, respectively.
Results
Molecular imprinting of tocopherols
Molecularly imprinted polymers toward TP, TPN and TPS have been successfully prepared. Results indicated that TP-MIP, TPN-MIP and TPS-MIP in DCM displayed very low binding capacity toward their respective template analytes. Such findings were in accordance with the results from our previous investigations (Piacham et al., 2009[51]). In order to resolve this situation, appropriate solvents for template rebinding were investigated using various amounts of aqueous-ethanol mixtures. It was found that the optimum ratio for the ethanol-aqueous mixture was 3:2, v/v. Such aqueous mixtures ratio displayed good rebinding and selective recognition for TP and its analogs as compared to the control non-imprinting polymer (NIP) as shown in Figure 2(Fig. 2). It should be noted that these MIPs were synthesized via thermal-induced polymerization. TP-MIP displayed recognition property greater than the MIP prepared from the previous report (Piacham et al., 2009[51]), which was also synthesized by thermal-induced polymerization. binding properties may be attributed to differences in the template and monomer content ratio and the porogenic solvent used (Puoci et al., 2007[58]). Particularly, results revealed that TP-MIP, TPS-MIP and TPN-MIP all bound its respective template molecules with 2-fold greater binding than the NIP at 10 mg polymer. The binding capacity for all MIPs was approximately 2 mg/g of polymer.
It was observed that NIP could bind TP and TPN rather non-specifically even when the polymer concentration was increased. On the other hand, NIP was shown to have low binding capacity towards TPS. It can be seen from Figure 2(Fig. 2) that 80 mg of TPS-MIP exhibited high affinity towards TPS as deduced from a binding capacity of greater than 92 % in comparison to 13 % binding by NIP at template concentration of 0.1 mg/mL while TPN-MIP, TPS-NIP, TP-MIP and TP-NIP provided binding performances of 78.2 %, 55.2 %, 92 % and 55 %, respectively. It can be seen that TPS-MIP provided over 7 folds higher % binding when compared to their control NIP while TPN-MIP and TP-MIP afforded roughly 1.5-folds higher % binding than the NIP. Such specific binding of TPS may be attributed to their inherent physicochemical properties in which the succinic moiety of TPS possesses both hydrogen bond donating and accepting properties. This may consequently lead to strong binding of TPS to MAA, which also contain hydrogen bond donating and accepting capacities. Such strong template-monomer interaction of TPS and MAA may potentially give rise to a rather defined binding cavity. On the other hand, the rather non-specific binding of MIP and NIP to their respective templates TP and TPN may be attributed to the long chain aliphatic moiety as well as the one-point monomer interactions at hydroxyl group of TP and pyridine nitrogen of TPN.
In light of the non-specific binding properties as afforded by TP-MIP and TPN-MIP, a second round of polymer synthesis was performed to produce molecularly imprinted nanospheres toward TP and TPS using the precipitation polymerization technique. Sizes and morphology of the obtained nanospheres were determined by SEM (Figure 3(Fig. 3)) to have uniform shape and size in the range of 200-400 nm, which may be suitable for future drug delivery applications. The binding capacities of TP-MIN, TPS-MIN and NIN were investigated by means of batch analysis in acetonitrile.
It was observed that 80 mg of TPS-MIN and NIN afforded binding capacities of 47 % and 19 %, respectively, while TP-MIN and NIN had binding performances of 35 % and 22 %, respectively (Figure 4(Fig. 4)). Such results indicated that TPS-MIN could bind to its template more specific than that of TP-MIN. To further enhance the binding performance of TPS-MIN, the binding solvents were subjected to optimization studies in which the aqueous content in organic solvent was varied. As shown in Figure 5(Fig. 5), 40 mg of TPS-MIN were incubated with 0.1 mg/mL of TPS in different solvent mixtures including EtOH:H2O (4:1, v/v), EtOH:H2O (3:2, v/v) and MeCN:H2O (1:1, v/v), which afforded % binding of 43 %, 22 % and 17 %, respectively for TPS-MIN while % binding of 10 %, 7 % and 8 %, respectively, for TPS-NIN. The results indicated that EtOH:H2O using a ratio of 4:1, v/v augmented the binding specificity of TPS-MIN by 4-folds higher than the control NIN.
Computer simulation of tocopherol-imprinted polymers
Mechanistic insights into the origin of the observed binding specificities for the tocopherol-imprinted polymers were deduced from computational chemistry calculations. Computational chemistry had been demonstrated to be useful in elucidating the physicochemical properties of chemical entities that is pertinent for understanding the origins of the investigated biological activities and chemical properties (Nantasenamat et al., 2012[40]; Piacham et al., 2006[49]; Prachayasittikul et al., 2007[56], 2010[55]; Suksrichavalit et al., 2008[65], 2009[64]).
First, the template and functional monomer were subjected to an initial geometry optimization at AM1 level followed by a refined optimization at DFT level using B3LYP/6-31G(d) and finally subjected to a single point energy calculation at B3LYP/6-311++G(d,p) level. Second, the optimized structures of template and functional monomer were used for subsequent geometry optimizations of template-monomer complexes by iteratively placing the functional monomer at each of the possible functional moiety of the template molecule. This resulted in several possible complexes and their corresponding interaction energy, which were obtained by calculating the energetic difference of the template, functional monomer and their complexes.
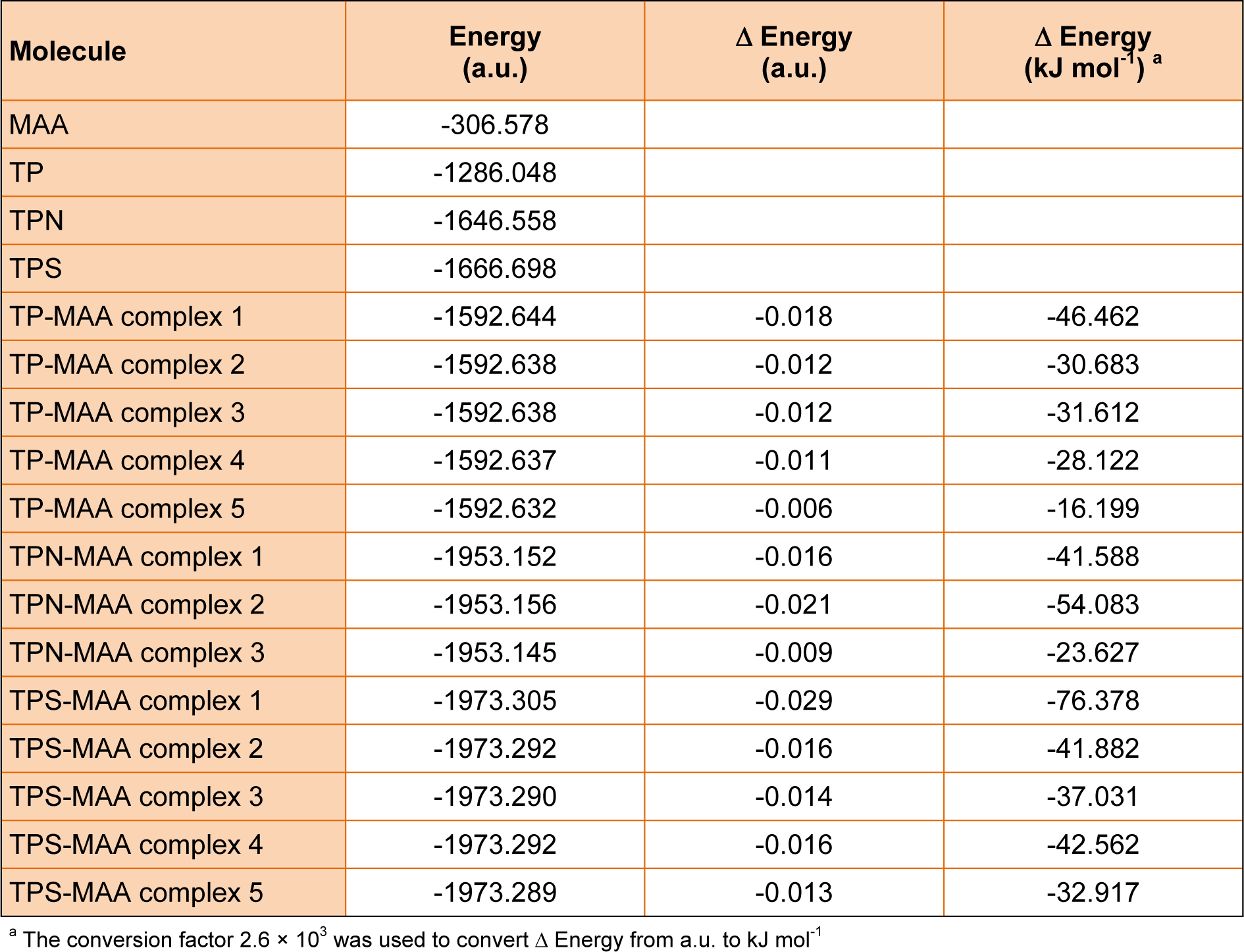
It can be seen from Table 1(Tab. 1) that TP-MAA complexes revealed interaction energies (in order of increasing values) of -46.462, -31.612, -30.683, -28.122 and -16.199 kJ mol-1 for TP-MAA complexes 1, 3, 2, 4 and 5, respectively, where its binding modalities are correspondingly shown in Figures 6a(Fig. 6), 6c(Fig. 6), 6b(Fig. 6), 6d(Fig. 6) and 6e(Fig. 6), respectively. It was observed from TP-MAA complex 1 that the hydroxyl group emanating from the benzopyran core structure acted as both hydrogen bond donor and acceptor by interacting with the two complementary functional moieties (i.e. hydroxyl and carbonyl groups) of MAA. Such two-point interaction with MAA accounted for the high interaction energy of TP-MAA complex 1. The remaining complexes essentially provided one-point interaction and consequently had lower interaction energy than that of TP-MAA complex 1.
The interaction energies for TPN-MAA complexes were -54.083, -41.588 and -23.627 kJ mol-1 for TPN-MAA complexes 2, 1 and 3, respectively, and its structures are correspondingly depicted in Figures 7b(Fig. 7), 7a(Fig. 7) and 7c(Fig. 7), respectively. As TPN could only provide hydrogen bond accepting capacity, its interaction with MAA were essentially one-point interactions. It was observed from TPN-MAA complex 2 that the nitrogen atom from the pyridine ring afforded the highest interaction energy while one-point interaction with oxygen atoms at various positions of TPN provided lower interaction energy.
Interaction energies of -76.378, -42.562, -41.882, -37.031 and -32.917 kJ mol-1 were observed for TPS-MAA complexes 1, 4, 2, 3 and 5, respectively, and its structures are correspondingly shown in Figures 8a(Fig. 8), 8d(Fig. 8), 8b(Fig. 8), 8c(Fig. 8) and 8e(Fig. 8), respectively. TPS-MAA complex 1 was found to afford the highest interaction energy of -76.378 and this could be attributed to the two-point interaction of MAA with the succinate moiety of TPS in which the carboxylic group provided both hydrogen bond donating and accepting capacities. The bond distances of interacting atoms at the site of this two-point interaction were 1.67 and 1.68 Å. Such distances were found to be the least of the investigated conformers thereby corroborating the observed high interaction energy.
The computer simulation suggested that template-monomer complexes providing the strongest binding were TPS-MAA > TPN-MAA > TP-MAA where the highest interaction energy were -76.378, -54.083 and -46.462, respectively. The high interaction energy afforded by TPS-MAA complex is in good agreement with the experimental finding that TPS-MIP also provided the best binding performance of up to 7-folds higher than the control NIP for bulk monoliths while 3-folds higher binding than the control NIP were observed for the uniformly-sized TPS-MIN.
Conclusion
Everyday we are constantly bombarded by a wide range of environmental factors that predisposes us to the development of cancer. Tocopherol succinate is a vitamin E derivative that has been shown to possess promising anti-cancer activity. In this work, molecularly imprinted polymers with bind- ing specificity toward tocopherol and derivatives were prepared by bulk polymerization and precipitation polymerization. TPS-imprinted polymers prepared by both methods were found to have higher specificity and selectivity than the other derivatives as deduced from 7- and 3-folds higher % binding for bulk and uniformly-sized particles, respectively. Nanospheres prepared by precipitation polymerization afforded polymers with uniform size (~200-400 nm) and shape, which are suitable for further applications in drug delivery efforts.
References
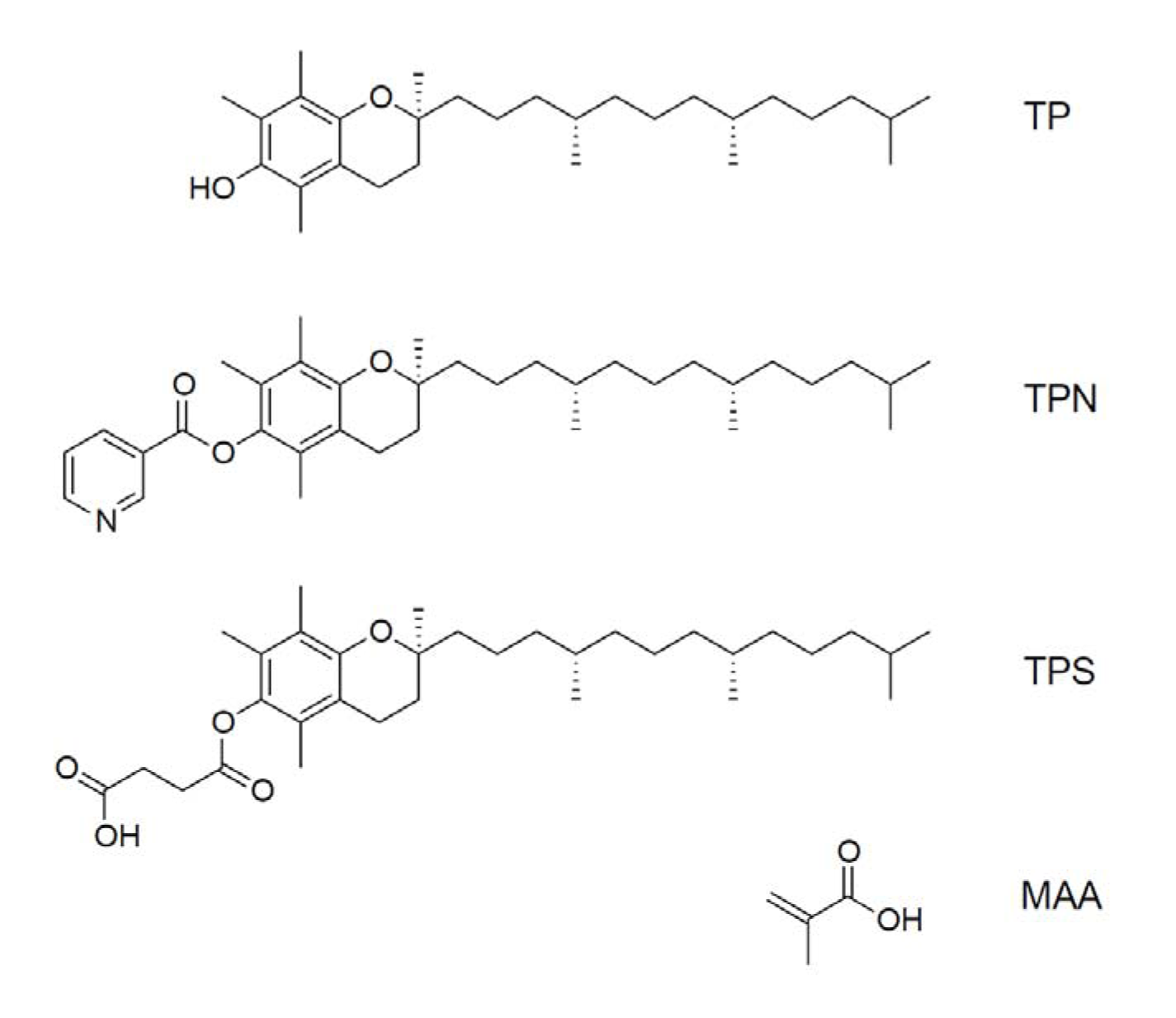
Figure 1: Chemical structures of tocopherol (TP), tocopherol nicotinate (TPN), tocopherol succinate (TPS) and methacrylic acid (MAA)
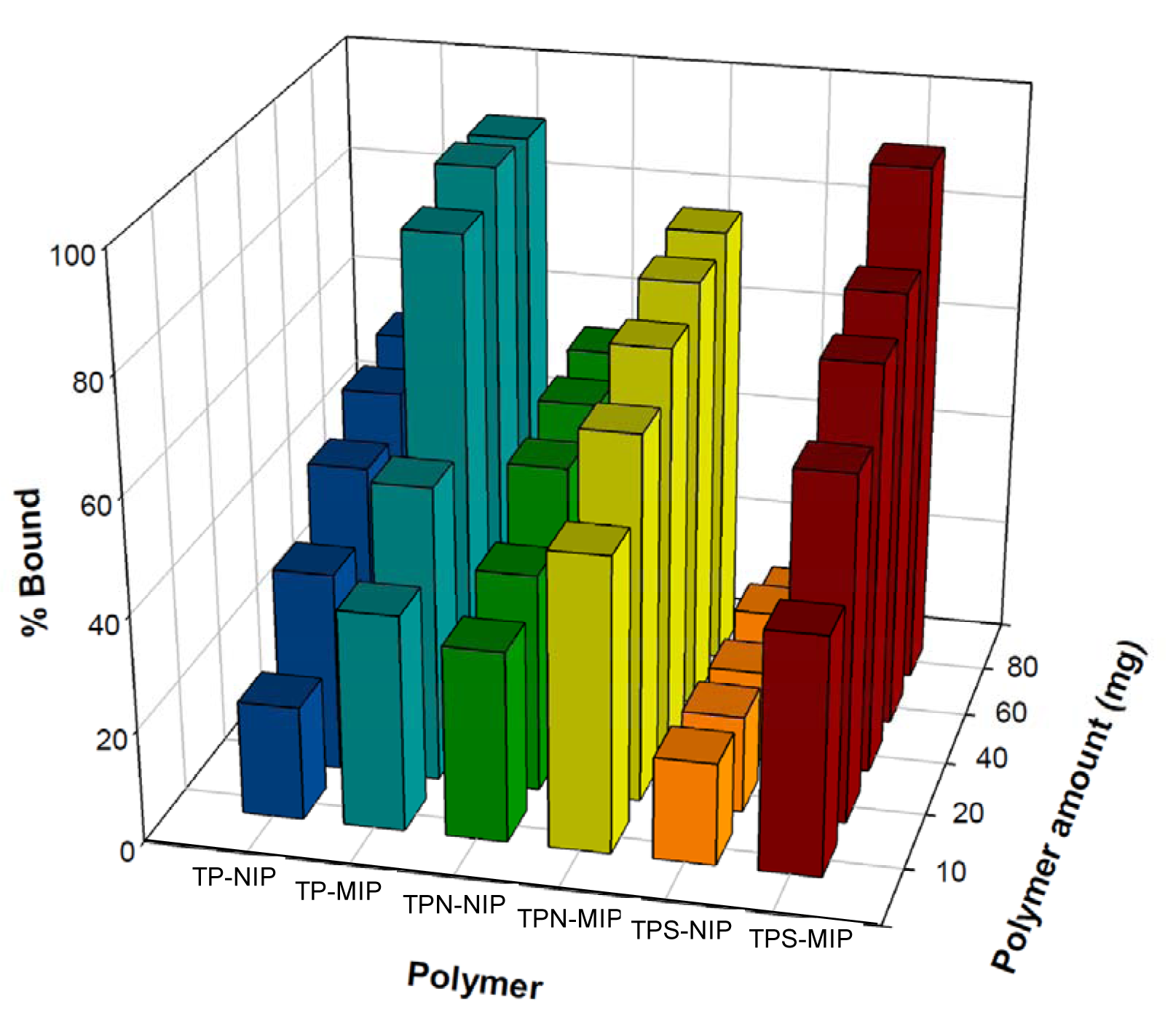
Figure 2: Rebinding experiment of tocopherol molecularly imprinted polymers (TP-MIP), tocopherol succinate molecularly imprinted polymer (TPS-MIP) and tocopherol nicotinate molecularly imprinted polymer (TPN-MIP) toward their template as well as their non-imprinted counterparts (TP-NIP, TPS-NIP and TPN-NIP)
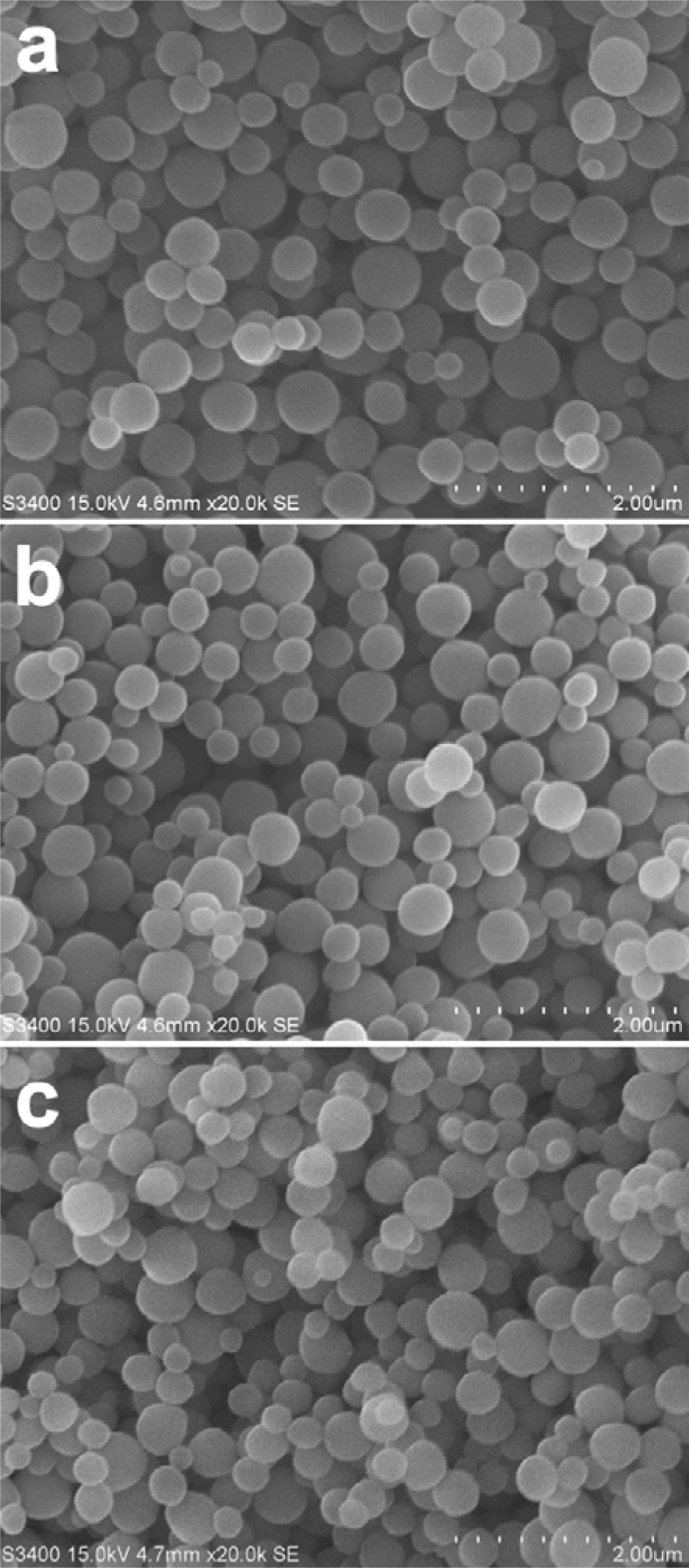

Figure 3: Scanning electron micrograph of topherol succinate molecularly imprinted nanospheres (TPS-MIN) (a), tocopherol molecularly imprinted nanospheres (TP-MIN) (b) and non-imprinted nanospheres (NIN) (c)
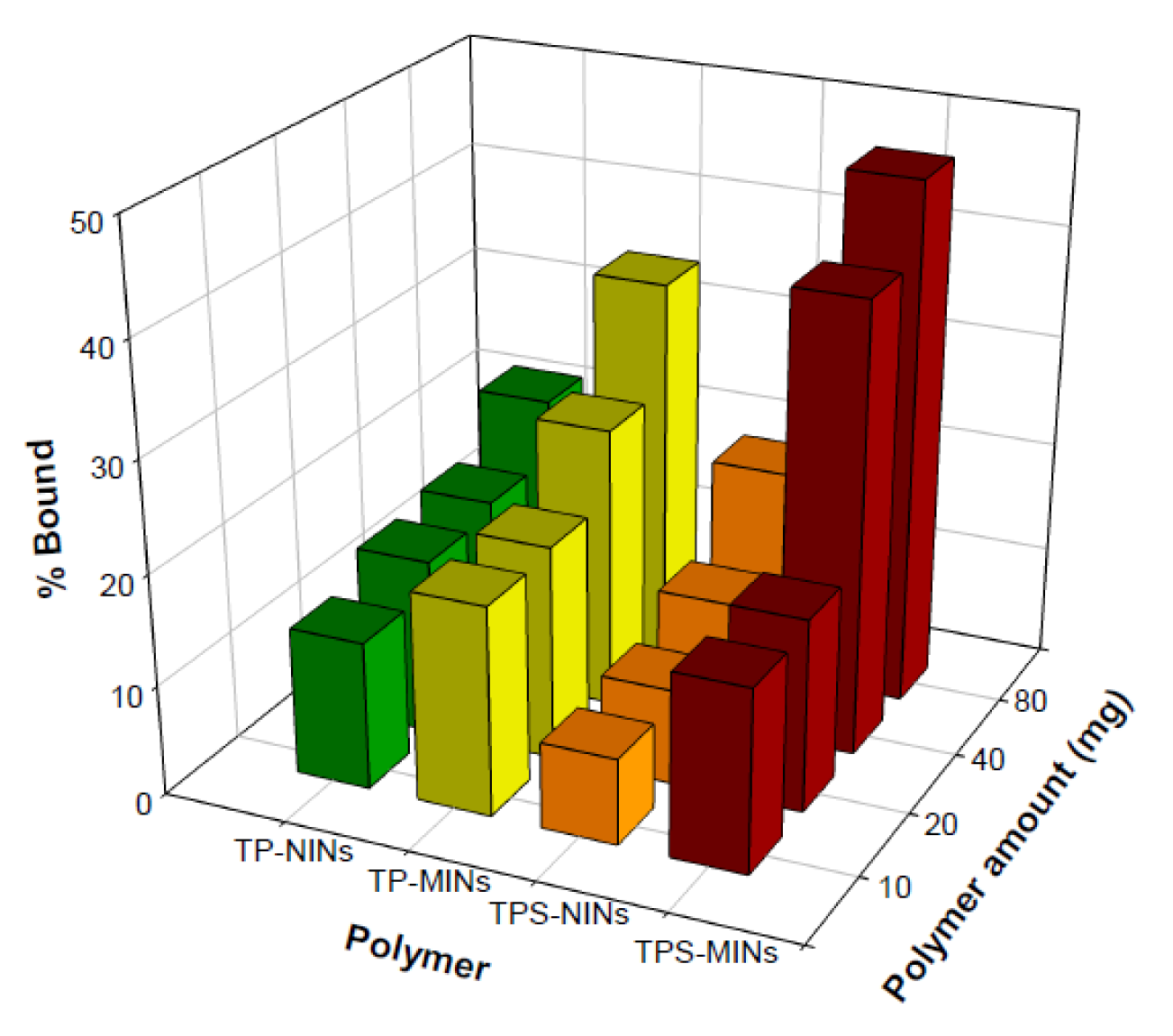
Figure 4: Rebinding experiment of tocopherol succinate molecularly imprinted nanospheres (TPS-MIN), tocopherol molecularly imprinted nanospheres (TP-MIN) and non-imprinted nanospheres (NIN) toward tocopherol succinate (TPS) and tocopherol (TP)
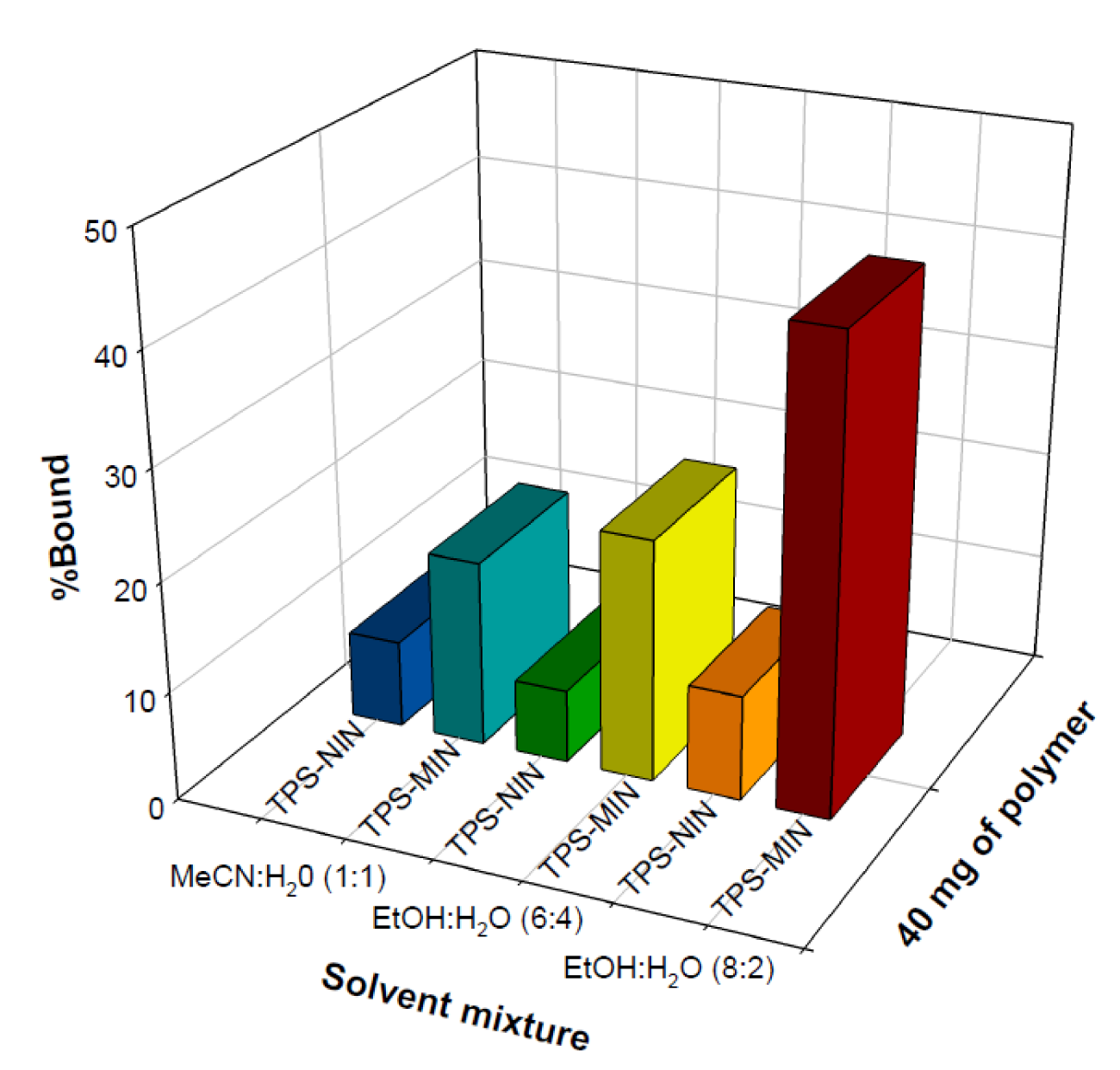
Figure 5: Optimization of binding solvent of tocopherol succinate molecularly imprinted nanospheres (TPS-MIN)
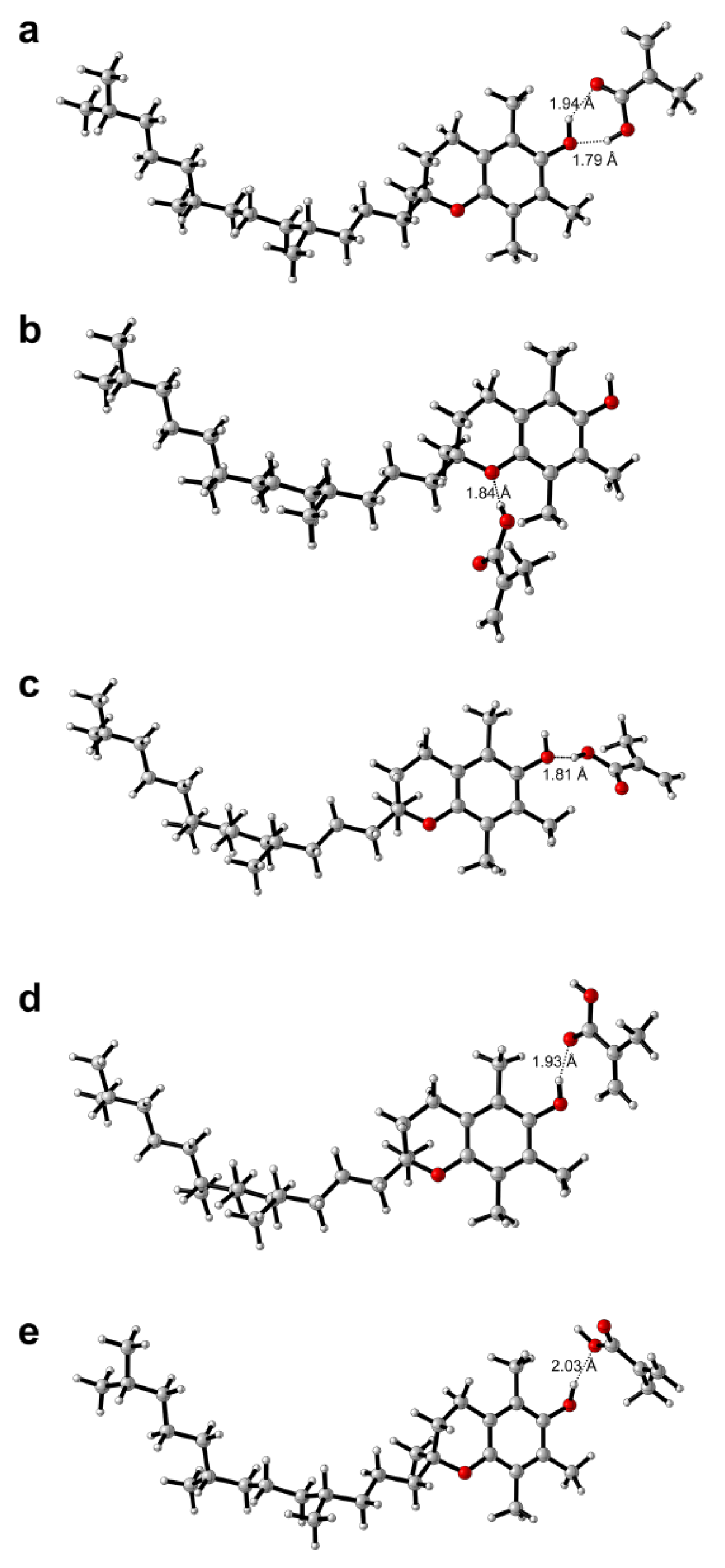
Figure 6: Possible binding modalities as obtained from computational chemistry calculations for tocopherol-methacrylic acid (TP-MAA) complexes 1-5 (a-e)
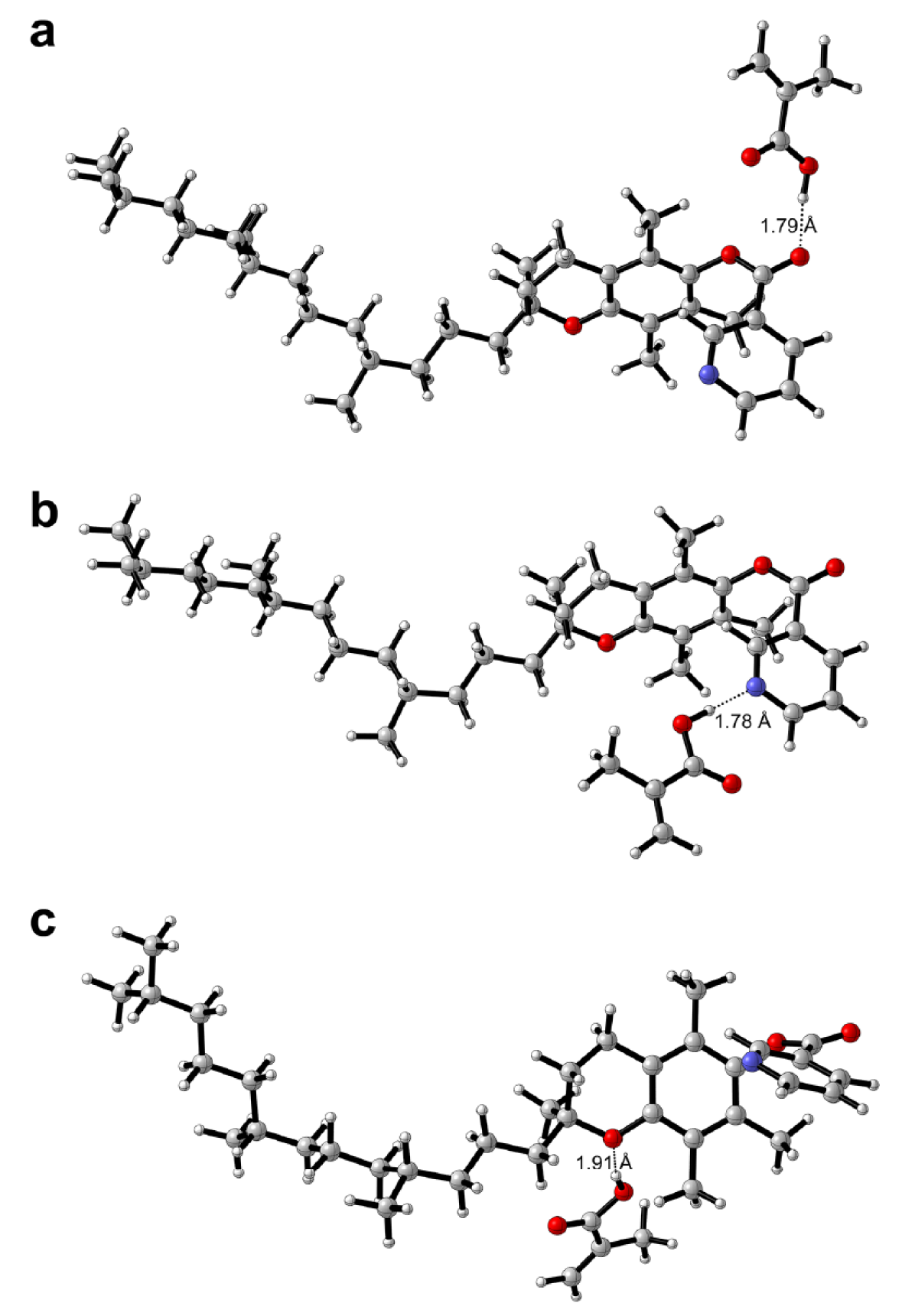
Figure 7: Possible binding modalities as obtained from computational chemistry calculations for tocopherol nicotinate-methacrylic acid (TPN-MAA) complexes 1-3 (a-c)
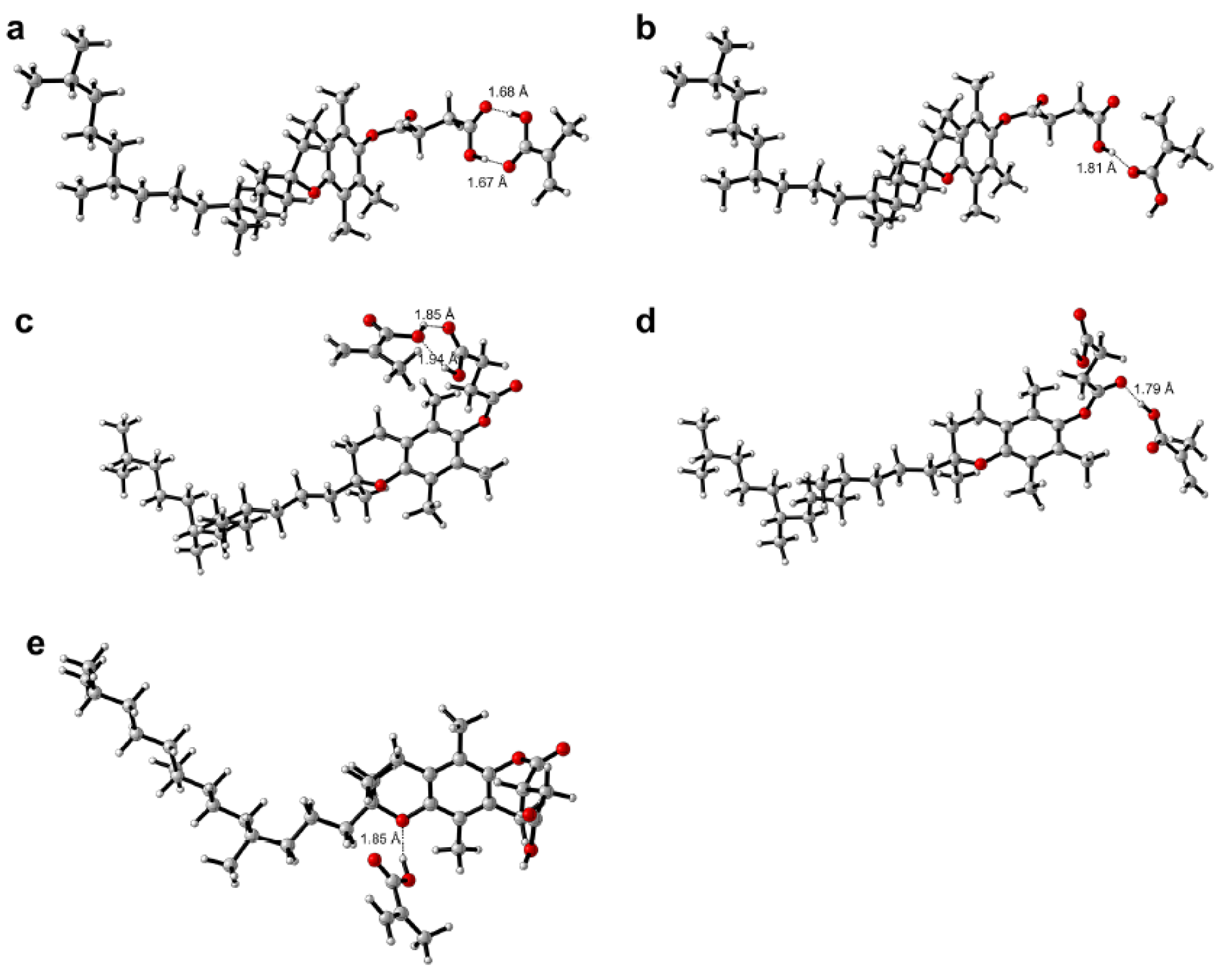
Figure 8: Possible binding modalities as obtained from computational chemistry calculations for tocopherol succinate-methacrylic acid (TPS-MAA) complexes 1-5 (a-e)
[*] Corresponding Author:
Theeraphon Piacham, Department of Clinical Microbiology and Applied Technology, Faculty of Medical Technology, Mahidol University, Bangkok 10700, Thailand; phone: +66 2 441 4371, fax: +66 2 441 4380, eMail: theeraphon.pia@mahidol.ac.th