Research article
Selenoprotein M is expressed during bone development
Melanie Grosch1, Jennifer Fuchs1, Michael Bösl2, Andreas Winterpacht1[*], Andreas Tagariello1
1Institute of Human Genetics, University Hospital Erlangen, University of Erlangen-Nürnberg, Erlangen, Germany2Max Planck Institute of Neurobiology, Martinsried, Germany
EXCLI J 2013;12:Doc967
Abstract
25 selenoproteins that contain selenium, incorporated as selenocysteine (Sec), have been identified to date. Selenoprotein M (SELM) is one of seven endoplasmic reticulum (ER)-resident, Sec-containing proteins that may be involved in posttranslational processing of proteins and maintenance of ER function. Since SELM was overrepresented in a cartilage- and bone-specific expressed sequence tag (EST) library, we further investigated the expression pattern of Selm and its possible biological function in the skeleton. RNA in situ hybridization of Selm in chicken and mice of different developmental stages revealed prominent expression in bones, specifically in osteoblast, and in tendons. This result suggests that SELM functions during bone development, where it is possibly involved in the processing of secreted proteins.
Keywords: selenium, selenoproteins, Kashin-Beck osteoarthropathy, bone development, thiol-disulfide oxidoreductase, unfolded protein response
Introduction
Selenium is an essential trace element that plays an important role in several metabolic pathways, including thyroid hormone metabolism, antioxidant defense system, and intracellular redox homeostasis (Gromer et al., 2005[11]). Moreover, selenium has been shown to be incorporated as selenocysteine (Sec) into at least 25 human and 24 rodent selenoproteins identified so far (Behne and Kyriakopoulos, 2001[1]; Kryukov et al., 2003[17]; Reeves and Hoffmann, 2009[24]). Sec is encoded by the UGA codon, and recognition of this 21st amino acid is ensured by a stem-loop structure in the 3′ UTR termed the Sec insertion sequence (SECIS) (Driscoll and Copeland, 2003[6]). Although their precise biological functions remain unknown, recent data suggest that endoplasmic reticulum (ER)-resident selenoproteins form an important part of the ER quality control machinery (Labunskyy et al., 2007[19]) and may be involved in the formation or isomerization of disulfide bonds and in the unfolded protein response (UPR) (Hawkes and Alkan, 2010[12]). The correct folding of proteins is essential for ER function and plays a fundamental role in the secretory pathway. Accumulation of misfolded or unfolded proteins in the ER results in ER stress, which leads to translational attenuation, transcriptional activation of chaperones, and protein degradation (Schroder and Kaufman, 2005[27]).
Selenoproteins are crucial for embryogenesis as targeted deletion of the Sec tRNA gene, Trsp, which is responsible for the expression of all selenoproteins, is embryonic lethal (Downey et al., 2009[5]). Furthermore, Col2a1 promoter-driven conditional knockout in osteochondroprogenitor cells revealed essential functions of selenoproteins in bone development. This corresponds to observations suggesting that bone tissue in man and rat is affected by selenium deficiency (Tsukahara et al., 1996[29]; Moreno-Reyes et al., 2001[22]). Moreover, selenium deficiency may also play a role in Kashin-Beck osteoarthropathy, endemic in China and Tibet (Moreno-Reyes et al., 1998[23]).
SELM is an ER-resident selenoprotein that is predicted to function as a thiol-disulfide oxidoreductase (Ferguson et al., 2006[8]). It contains a thioredoxin-like fold and a conserved CxxU active-site motif, which is characteristic of enzymes involved in the regulation of redox processes (Martin, 1995[21]). We recently carried out an expressed sequence tag (EST) project to identify genes and pathways involved in skeletal development. As a result, we observed that Selm was overrepresented in human growth plate tissue (Tagariello et al., 2005[28]). However, although SELM is thought to be involved in oxidative stress (Hwang et al., 2008[14]; Yim et al., 2009[30]; Garcia-Triana et al., 2010[10]) and Alzheimer's disease (Hwang et al., 2005[13]), the expression pattern and tissue-specific function of Selm remain unknown.
In the present study, we describe for the first time prominent Selm expression in bony structures and tendons of the developing chicken and mouse. The present data suggest that SELM may play a role in the proper folding and processing of secreted proteins during bone development.
Results and Discussion
We first analyzed the expression pattern of SELM by RNA in situ hybridization. Due to an easier handling and better availability (especially of early stages), tissue sections of chicken wings aged 6 days (Hamburger Hamilton stage 29; HH29) to 14 days (HH40) were used (Figure 1A(Fig. 1)). At HH29 we observed SELM expression in the condensed mesenchyme that will develop into bone. In later developmental stages prominent expression in bones could be detected. This could be verified by detailed comparative analysis of serial sections of wings HH40 with different markers (Figure 1B, C(Fig. 1)). The SELM expression pattern largely coincides with that of osteocalcin (BGLAP) and also with alkaline phosphatase (AP), two markers for osteoblasts (Ducy and Karsenty, 1995[7]; Schinke and Karsenty, 1999[25]; Franz-Odendaal et al., 2006[9]). These results suggest expression in osteoblasts. To exclude expression in chondrocytes and verify expression in osteoblasts, we carried out a semiquantitative PCR with cDNA of different cell types (Figure 2(Fig. 2)). In chicken micromass cultures, differentiated to chondrocytes for seven days, no SELM expression could be detected (Figure 2A(Fig. 2)). The chondrocyte phenotype was verified by type II (COL2A1) and type X (COL10A1) collagen, two markers for chondrocytes (Lefebvre and Smits 2005[20]). In contrast, expression of selm could be detected in primary osteoblasts prepared from chicken calvariae and differentiated for 21 days (Figure 2B(Fig. 2)). Also in the murine pre-osteoblastic cell line MC3T3 (differentiated for 30 days) Selm could be detected together with the osteoblast markers Osteocalcin (BGLAP) and type I collagen (COL1A1) (Figure 2C(Fig. 2)).
Additional to expression in osteoblasts, we also observed expression in tendons. By RNA in situ hybridization of wings of different developmental stages we detected coexpression of SELM, scleraxis (SCX), and tenomodulin (TNMD) (Figures 3(Fig. 3), 4(Fig. 4)). Since SCX and TNMD are specifically expressed in tenocytes (Blitz et al., 2009[2]), our results demonstrate expression of SELM in this cell type.
To verify and further support the strong expression pattern of SELM especially in the developing bone we used another species and subsequently analyzed Selm expression on murine tissue sections by RNA in situ hybridization. On embryonic day E16.5, Selm was prominently expressed in the bones of the craniofacial, axial, and appendicular skeleton (Figure 5(Fig. 5)). In particular, expression in the vertebrae, maxilla, and mandibula could be detected (Figure 5A(Fig. 5)) as well as in the trabecular bone and the periosteum of long bones (Figure 5B(Fig. 5) and data not shown). This expression pattern largely coincides with that of Bglap (Figure 5A(Fig. 5)). In addition, we also observed expression of Selm in the olfactory epithelium at this developmental stage (Figure 5A(Fig. 5)). We subsequently analyzed expression of Selm in the head and limb sections of 4-day-old mice (P4) (Figure 6(Fig. 6)). Strong expression of Selm was detected in the cerebellum and olfactory epithelium, and to a lesser extent in the hippocampus (Figure 6A(Fig. 6)). These results corroborate the recent findings of a study by Zhang et al. (2008[31]), which demonstrated elevated Selm expression in the cerebellum, olfactory bulb, Ammon's horn, and hippocampus. We found additional expression in the teeth and bones that resembled the expression pattern of Bglap (Figure 6A(Fig. 6)). Moreover, expression of Selm was observed in the periosteum and trabecular bone of P4 hindlimb sections (Figure 6B(Fig. 6)). Comparison of the expression pattern of Selm with that of markers for early (Col1a1) and late (Bglap) osteoblasts (Jikko et al., 1999[15]; Cohen, 2006[4]; Kulterer et al., 2007[18]), revealed overall coexpression of Selm and Col1a1. In contrast, coexpression of Selm and Bglap was restricted to the periosteum (Figure 6B(Fig. 6)).
Since strong Selm expression was observed in osteoblasts during bone development, we aimed to elucidate potential functions of SELM in osteoblasts in vivo. We generated a mutant mouse strain from ES cells (C57BL/6) containing a genetic deletion of Selm (Selm-/-; Supplementary Figure 1Winterpacht_Supplementary Figure 1.pdf). The open reading frame (ORF) of Selm in these ES cells was replaced with a ZEN-Ub1 cassette that contains a LacZ reporter gene under transcriptional control of the endogenous gene promoter. Although complete loss of Selm expression was verified by RNA in situ hybridization and real-time PCR (Supplementary Figure 1Winterpacht_Supplementary Figure 1.pdf), Selm deficiency did not affect viability or fertility. Moreover, morphological and histological analyses did not reveal any visible skeletal abnormalities, indicating that Selm deficiency may be compensated by other selenoproteins. Nevertheless, we used the Selm-/- mice to verify the expression pattern of Selm by X-Gal staining of E16.5 (Figure 7A(Fig. 7)) and P4 (Figure 7B(Fig. 7)) mice, which express β-galactosidase under the control of the Selm promoter (Supplementary Figure 1Winterpacht_Supplementary Figure 1.pdf). As shown in Figure 7A (e, f)(Fig. 7) comparative Alizarin red and Alcian blue staining for visualization of bone and cartilage, respectively, confirmed expression of Selm in almost all bony structures of the skeleton.
The present study is the first to analyze SELM expression in chicken and mouse sections during development and to link SELM with bone and tendon development. The ER-localized SELM protein possesses (1) a thioredoxin-like domain, (2) a surface-accessible active-site redox motif, and (3) structural similarities to other characterized thiol-disulfide oxidoreductases (Korotkov et al., 2002[16]), suggesting that SELM represents a thiol-disulfide isomerase that is involved in disulfide bond formation in the ER (Ferguson et al., 2006[8]). Moreover, the high expression level of SELM in osteoblasts and tenocytes - the main matrix producing cells of bones and tendons, respectively - indicates that SELM plays a role in the processing of ECM proteins. The ER is the site of synthesis and processing of secretory pathway proteins, which include transmembrane proteins and constituents of the extracellular matrix. In osteoblasts and tenocytes, vast amounts of ECM proteins, particularly the fibrillar collagens COL1A1 and COL1A2, are synthesized and secreted. Correct folding is a prerequisite for exiting the ER, whereas misfolded proteins are retained. Accumulation of misfolded or underglycosylated proteins results in ER stress - a process that has been shown to play a fundamental role in the initiation and progression of a wide range of connective tissue disorders (Boot-Handford and Briggs 2010[3]). Together with observations demonstrating an involvement of SELM in unfolded protein response (UPR) and ER stress (Grosch et al., unpublished), the data indicate that SELM may play a role in the proper folding and processing of secreted proteins during bone and tendon development.
Experimental Procedures
Ethics statement
All mouse experiments were approved by the Ethics Committee for Animal Experiments of the University of Erlangen (TS-05/10).
RNA in situ hybridization
RNA in situ hybridization on 5 µm paraffin sections of chicken or mouse tissue using digoxigenin-labeled antisense riboprobes for Selm, Col1a1, and Bglap was carried out as previously reported (Schmidl et al., 2006[26]). Specific cDNA fragments for murine Selm (NM_053267) and Bglap (NM_001032298) antisense riboprobes were obtained by RT-PCR
- (Selm, 5′-ttcttgcagcccttgtggctcc-3′ and 5′-acttgcggtagaagccgagc-3′;
- Bglap, 5′-catgaggaccctctctctgc-3′ and 5′-tgccagagtttggctttagg-3′) and cloned into pCRII-TOPO and pCR4-TOPO vectors, respectively. Specific cDNA fragments for chicken SELM, BGLAP (NM_205387), SCX (NM_204253) and TNMD (NM_206985) antisense riboprobes were obtained by RT-PCR
- (SELM, 5′-gcggtgggtgacggctgagccgg-3′ and 5′-ttacaggtctgggtggtccttcttg-3′;
- BGLAP, 5′-gctcacattcagcctctgc-3′ and 5′-ggagaagtggagcataatgg-3′;
- SCX, 5′-agctgtccaagatcgagacg-3′ and 5′-gtgaagacgggagtgtgtcc-3′;
- TNMD, 5′-accttcaggagtgggaacg-3′ and 5′-tccgtgtagtcgttgacagg-3′) and cloned into the pCR4-TOPO vector. The Col1a1 plasmid was a kind gift from Dr. Michael Stock.
Cell culture
The pre-osteoblast murine cell line MC3T3 (ATCC) was cultivated in alpha modified Eagle's medium supplemented with 10 % FCS and 100 U/ml penicillin/ streptomycin at 37 °C and 5 % CO2. Cells were differentiated for 30 days by adding of 50 mg/ml ascorbic acid and 50 mg/ml glycerol-2-phosphate to the culture medium.
Micromass cultures
Micromass cultures were prepared from chicken limb buds (HH22-24) by digestion with 3 mg/ml dispase followed by treatment with 0.1 % collagenase Ia and 0.1 % trypsin. Cells were cultivated in DMEM:F-12 (1:1) supplemented with 10 % FCS, 0.2 % CS and 100 U/ml penicillin/streptomycin at 37 °C and 5 % CO2. Culture medium was changed every day.
Primary osteoblasts
Primary osteoblasts were prepared from the calvariae of chicken (HH41). Chicken calvariae were digested with 1,5 U/ml collagenase P and 0,05 % trypsin.
Primary osteoblasts were cultivated in alpha modified Eagle's medium supplemented with 10 % fetal calf serum (FCS) and 100 U/ml penicillin/streptomycin in 6-well plates at 37 °C and 5 % CO2. The culture medium was changed every 3 days. When the cells reached confluency, they were differentiated for 21 days by adding of 50 mg/ml ascorbic acid and 50 mg/ml glycerol-2-phosphate to the culture medium.
RT-PCR
Total RNA was isolated from cells using TRIzol (Invitrogen). First-strand cDNA was synthesized with oligo(dT) primer and Superscript III Reverse Transcriptase (Invitrogen), according to manufacturer's instructions.
To verify the osteoblast or chondrocyte phenotype of cell cultures, semiquantitative RT-PCR with following primers was carried out
- (COL1A2, 5′-gcaacattggattccctggacc-3′ and 5′-gttcacccttttctcccttgcc-3′;
- COL2A1, 5′-ccaacaccgccagcatcc-3′ and 5′-gccaatatccacgccaaactcct-3′;
- COL10A1, 5′-gccaatatccacgccaaactcct-3′ and 5′-cagaggaatagagaccat-3′;
- COL12A1, 5′-gcagaaccaaacctctcact-3′ and 5′-ttcttggtgttcctctctcc-3′;
- SELM, 5′-gcggtgggtgacggctgagccgg-3′ and 5′-ttacaggtctgggtggtccttcttg-3′;
- Bglap, 5′-catgaggaccctctctctgc-3′ and 5′-tgccagagtttggctttagg-3′;
- Col1a1, 5′-cctggtaaagatggtgcc-3′ and 5′-caccaggttcacctttcgcacc-3′;
- Selm, 5′-ttcttgcagcccttgtggctcc-3′ and 5′-acttgcggtagaagccgagc-3′).
- Gapdh was used as control (GAPDH, 5′-gtggagtccactggtgtcttc-3′ and 5′-atcagcagcagccttcactac-3′;
- Gapdh, 5′-gtggagtccactggcgtcttc-3′ and 5′-ctccgacgcctgcttcaccac-3′).
Alcian blue and Alizarin red staining
Whole E15.5 embryos or skinned and eviscerated P4 mice were fixed in 95 % EtOH for 1-2 days. Mice were moved into glass vials and immersed in acetone overnight at room temperature. Subsequently, mice were rinsed in 95 % EtOH and stained for cartilage in Alcian blue (0.015 % Alcian blue 8GX, 76 % EtOH, and 20 % glacial acid). After washing with 95 % EtOH overnight and clearing in 1 % KOH (embryo) or 2 % KOH (P4), mice were stained in 0.005 % Alizarin red S in 1 % KOH for 3 to 6 h. For clearing of stained mice, samples were incubated in 5 % KOH for 5 days followed by a series of clearing solutions containing increasing glycerol and decreasing KOH concentrations (80:20, 60:40, 40:60, 20:80; 1 % KOH:glycerol). Finally, samples were stored in 100 % glycerol.
X-Gal staining
Whole E15.5 embryos or skinned and eviscerated P4 mice were fixed in 4 % paraformaldehyde for 1 h at 4°C. After 3 washing steps in pre-washing solution (1× PBS and 2 mM MgCl2) at 4°C, samples were incubated twice for 15 min at room temperature (RT) in detergent solution (1× PBS, 2 mM MgCl2, 0.02 % NP-40, and 0.01 % Na-deoxycholate) before incubation at RT with staining solution (1× PBS, 20 mM Tris-HCl [pH 7.5], 2 mM MgCl2, 0.02 % NP-40, 0.01 % Na-deoxycholate, 5 mM K3Fe(CN)6, 5 mM K4Fe(CN)6, and 0.75 mg/ml X-Gal in DMSO) under visual observation. Subsequently, samples were washed 3 times in post-washing solution (1× PBS and 5 mM EDTA, pH 8) and post-fixed overnight at 4°C in 4 % paraformaldehyde. For clearing of X-Gal stained mice, samples were incubated in 5 % KOH for 5 days followed by a series of clearing solutions containing increasing glycerol and decreasing KOH concentrations (80:20, 60:40, 40:60, 20:80; 1 % KOH:glycerol). Finally, samples were stored in 100 % glycerol.
Acknowledgements
We would like to thank Dr. Michael Stock for providing the Col1a1 plasmid.
References
File-Attachments
- Winterpacht_Supplementary Figure 1.pdf (101.05 KB)
Supplementary Figure 1
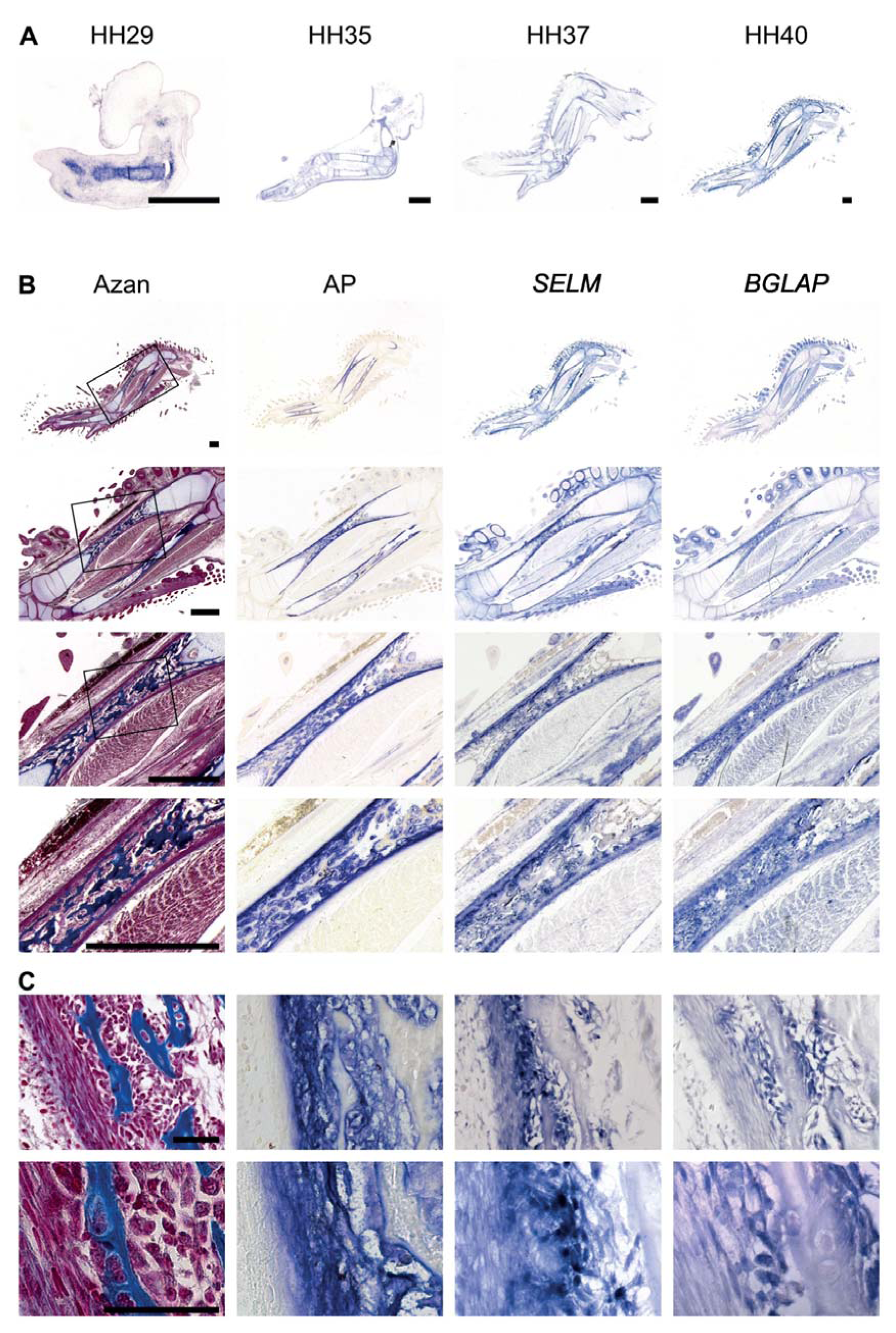
Figure 1: SELM expression in long bones during chicken development
RNA in situ hybridization on 5 µm thick paraffin sections of wings of indicated developmental stages (A) and wings at stage HH40 (B).
(A) Sections were hybridized with antisense riboprobes against SELM. Scale bar, 1 mm.
(B, C) Serial sections were hybridized with antisense riboprobes against SELM and BGLAP, an additional Azan and alkaline phosphatase (AP) staining is shown. Scale bar, 1 mm (B), 50 µm (C).
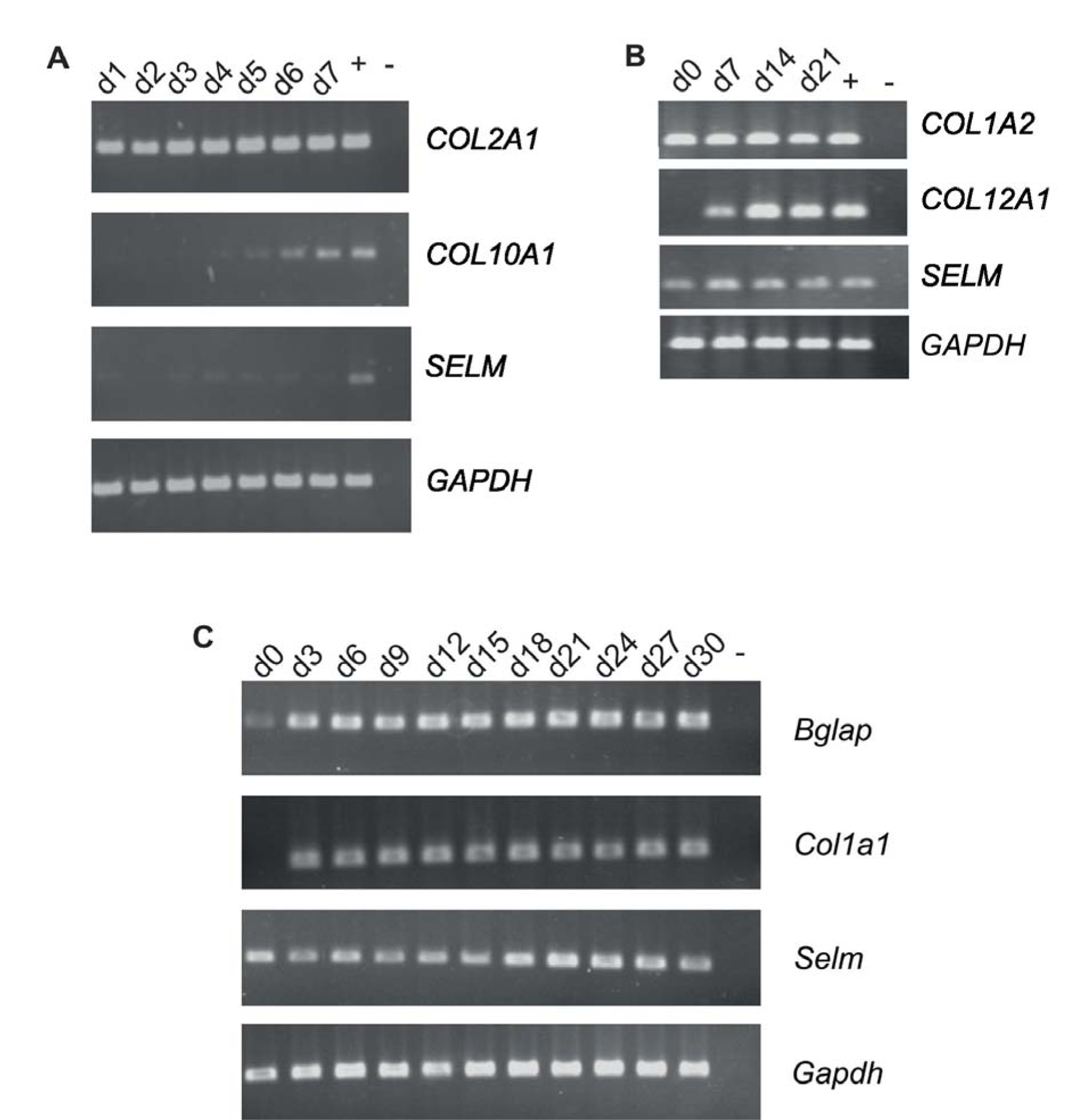
Figure 2: SELM expression in osteoblasts
Semiquantitative RT-PCR analysis of SELM expression in chicken micromass cultures (A), primary osteoblasts prepared from chicken calvariae HH41 (B) and in the murine pre-osteoblast cell line MC3T3 (C). Additional genes (as indicated) were used to verify the cell type, Gapdh served as loading control.
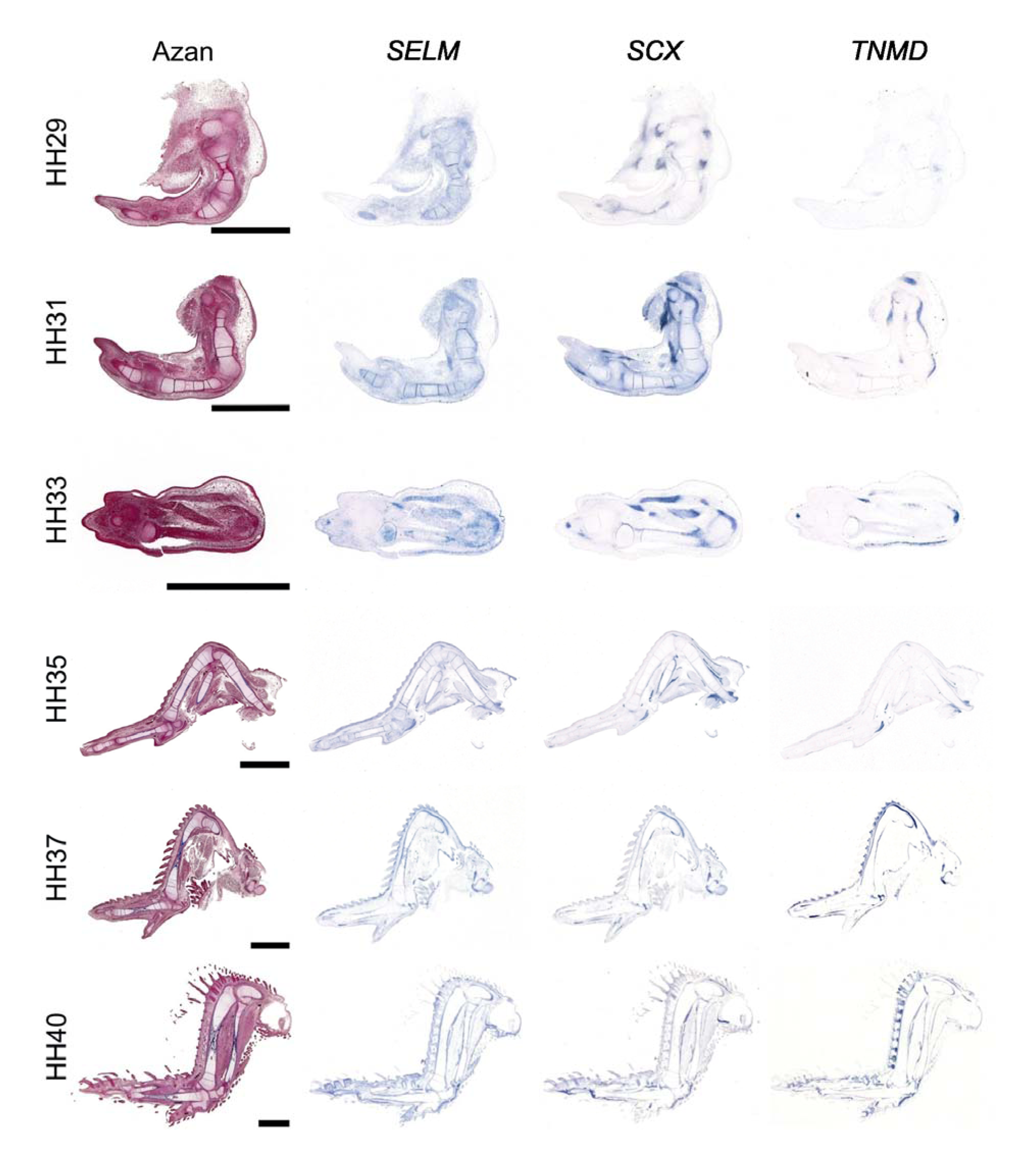
Figure 3: Expression of SELM in tendons during chicken development
RNA in situ hybridization on 5 µm thick paraffin sections of wings of indicated developmental stages. Serial sections were hybridized with antisense riboprobes against SELM, SCX and TNMD, an additional Azan staining is shown. Scale bar, 2 mm.
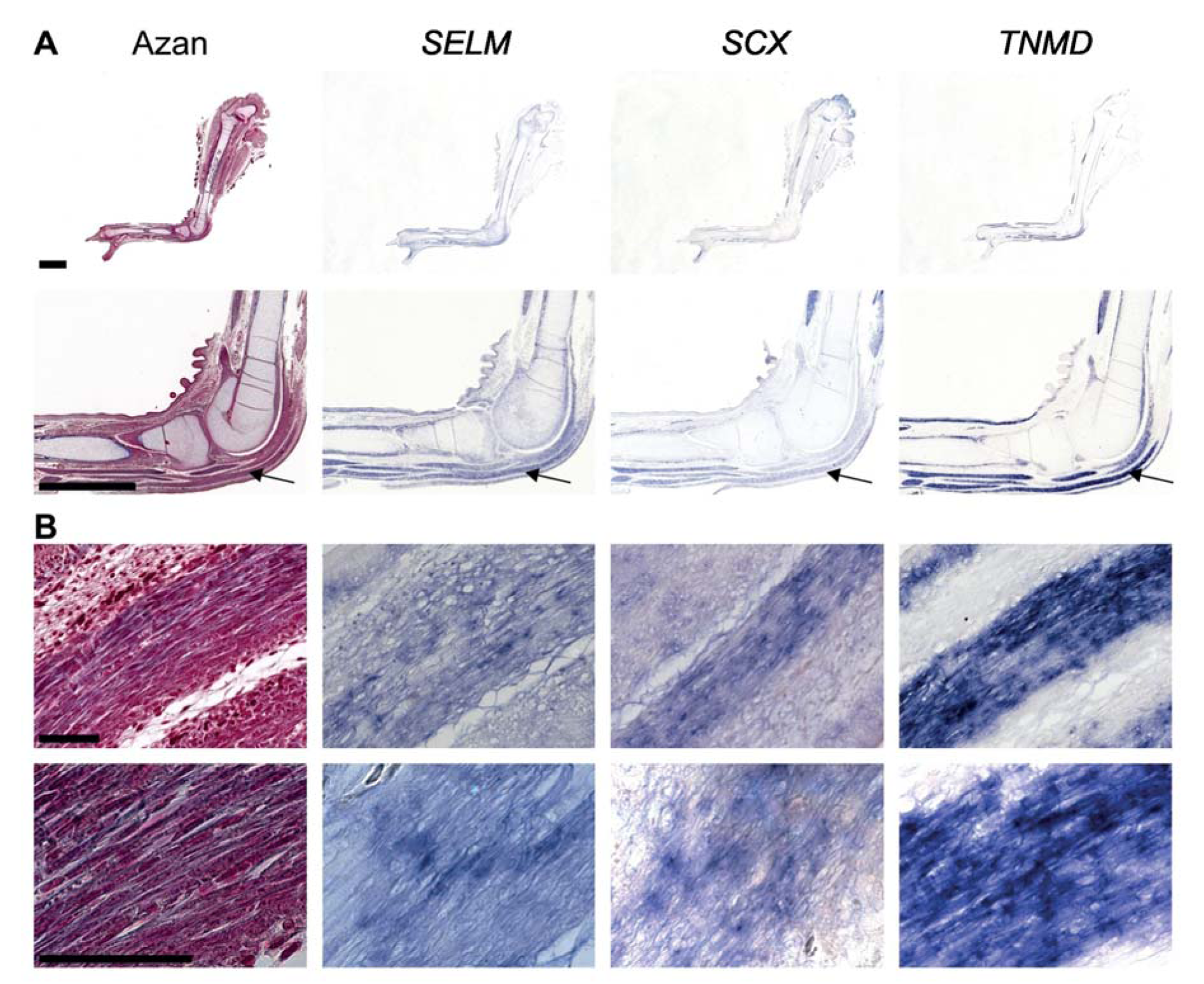
Figure 4: SELM expression in tenocytes
RNA in situ hybridization on 5 µm thick paraffin sections of legs at stage HH38. Serial sections were hybridized with antisense riboprobes against SELM, SCX and TNMD, an additional Azan staining is shown. One tendon is marked by an arrow. Scale bar, 2 mm (A), 50 µm (B).
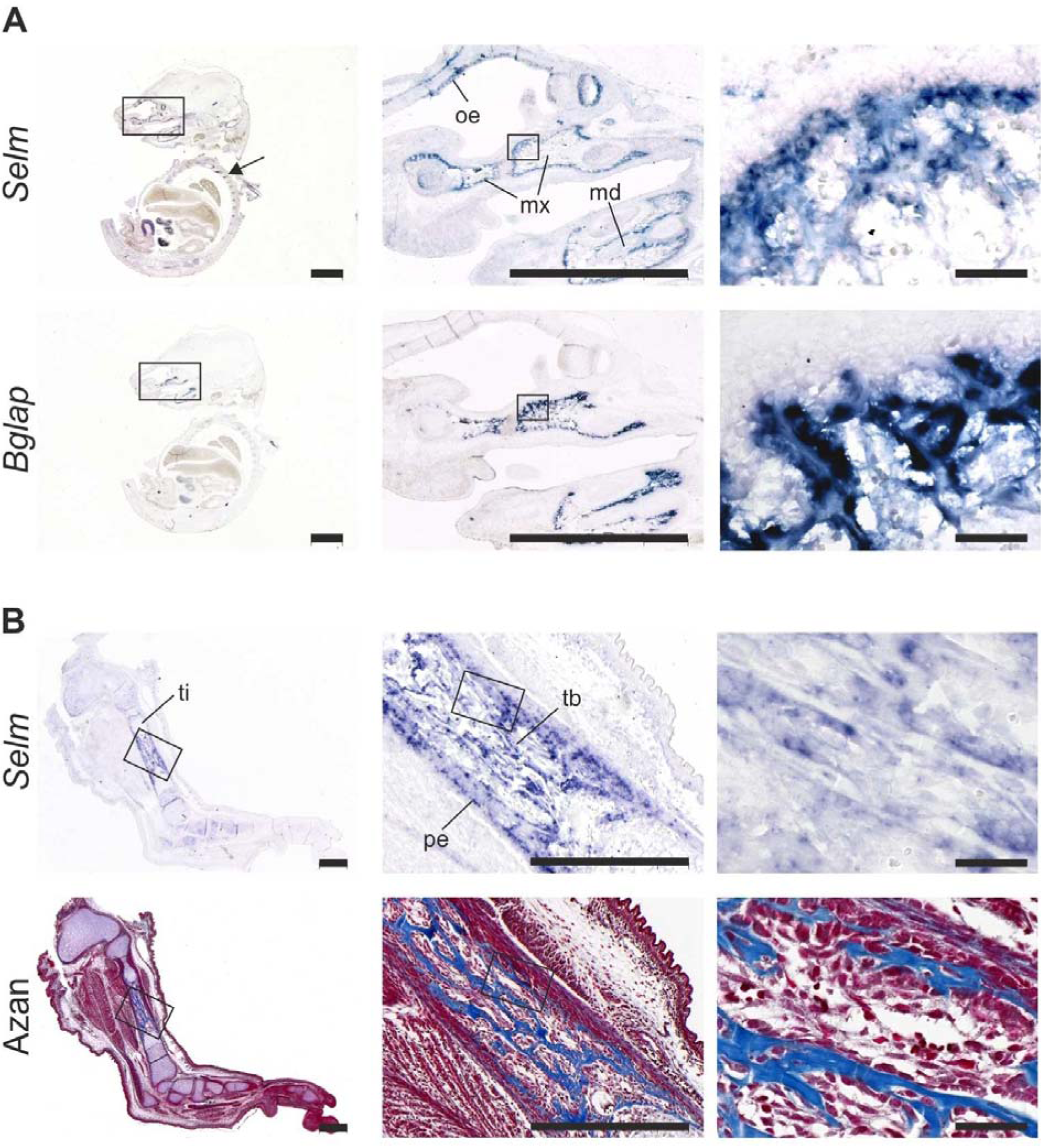
Figure 5: Expression of Selm in fetal mice (E16.5)
RNA in situ hybridization on 5 µm thick paraffin sections of whole embryos (A) and hindlimbs (B). Scale bar, 2 mm.
(A) Sections were hybridized with antisense riboprobes against Selm and Bglap as indicated. Vertebrae are marked by an arrow. md, mandibula; mx, maxilla; oe, olfactory epithelium.
(B) Sections were hybridized with antisense riboprobes against Selm, an additional Azan staining is shown. Pe, periosteum; tb, trabecular bone; ti, tibia.
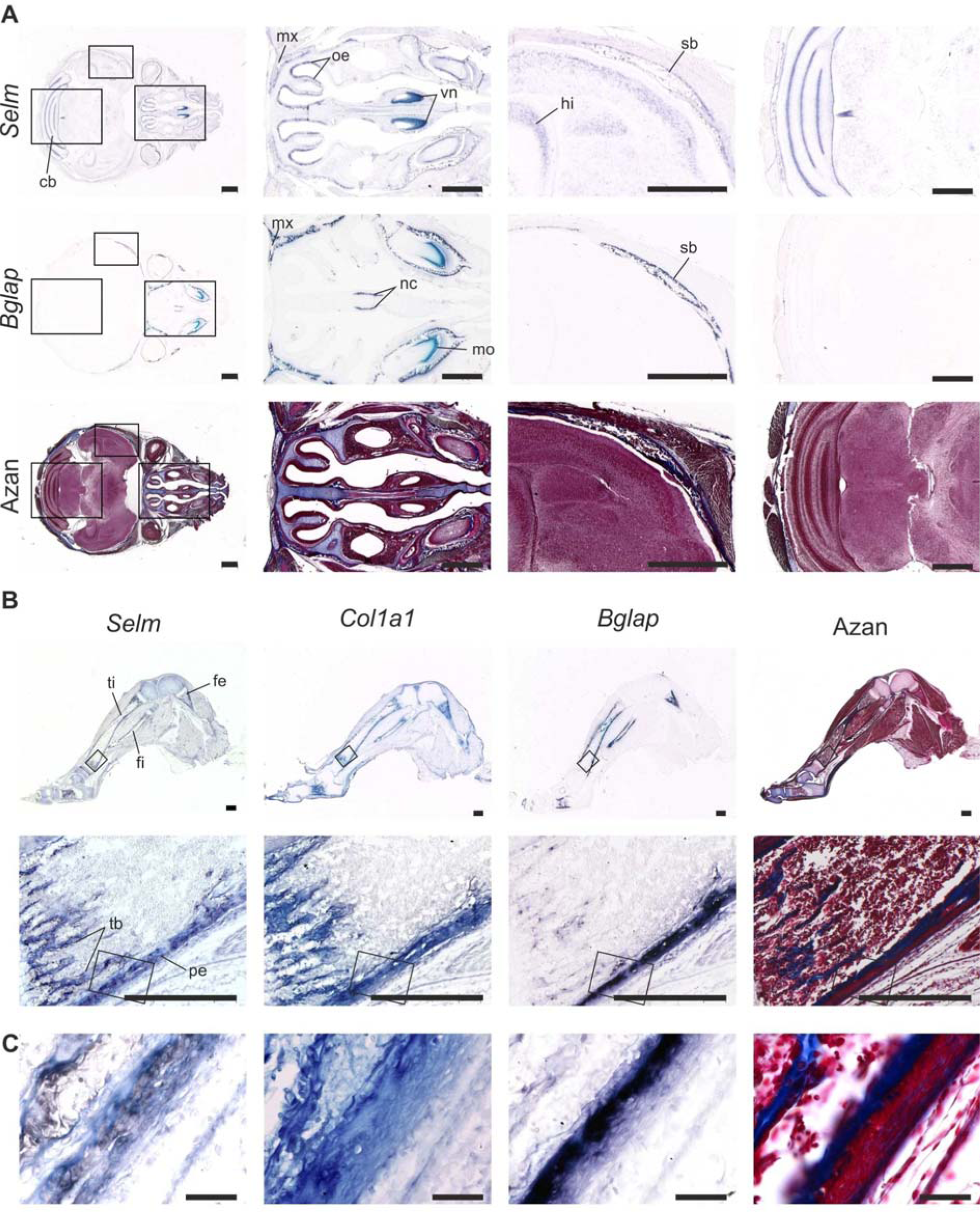
Figure 6: Expression of Selm in juvenile mice (P4)
RNA in situ hybridization on 5 µm thick paraffin serial sections of the head of postnatal day 4 (P4) mice (A; scale bar, 1 mm) and hindlimbs (B; scale bar, 0.5 mm). Sections were hybridized with antisense riboprobes against Selm, Bglap, and Col1a1, and stained with Azan as indicated. (C) Magnified views of the corresponding boxed areas in B. cb, cerebellum; fe, femur; fi, fibula; hi, hippocampus; mo, molar; mx, maxilla; nc, nasal concha; oe, olfactory epithelium; sb, sphenoid bone; tb, trabecular bone, ti, tibia; pe, periosteum; vn, vomeronasal organ.
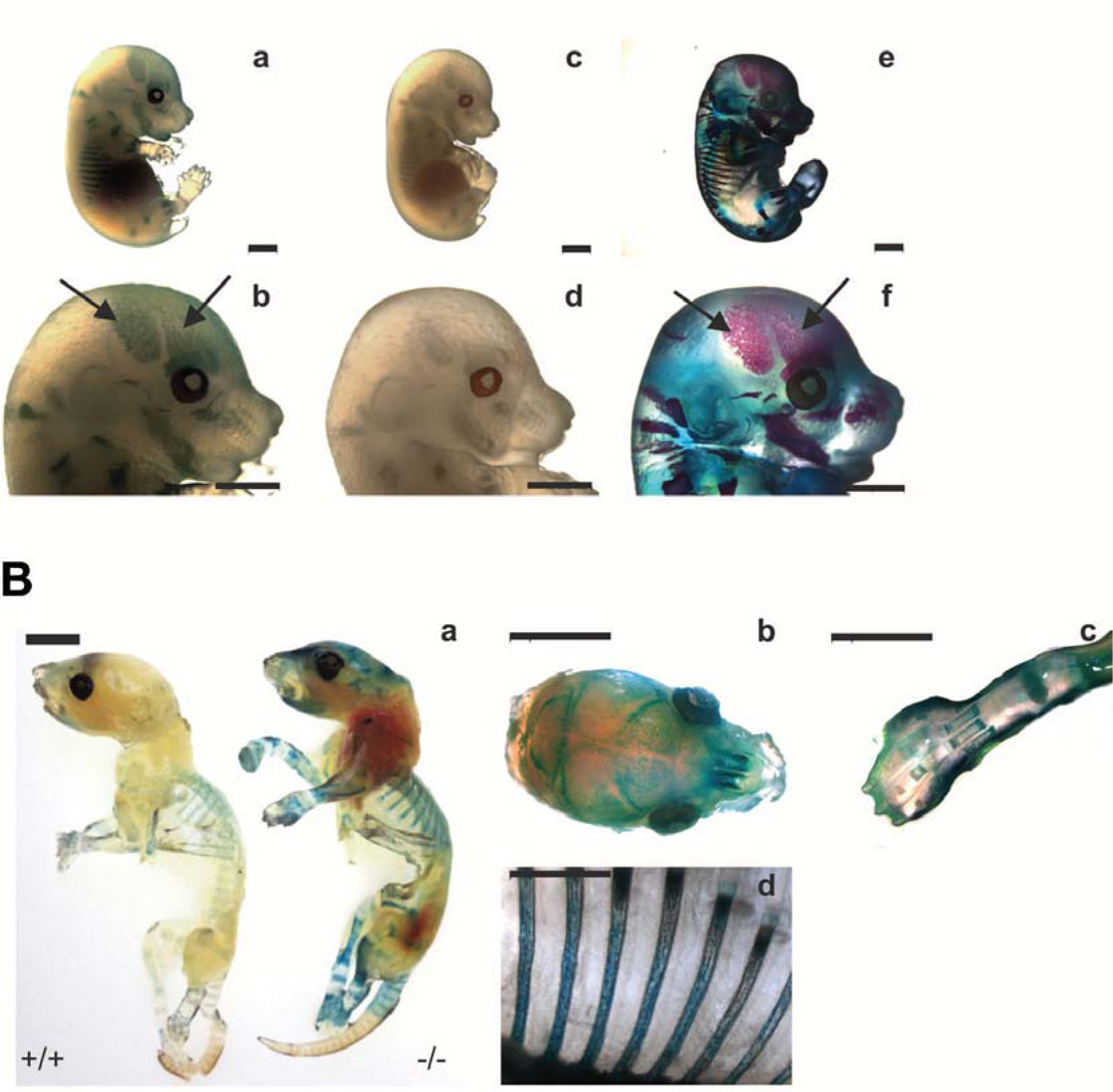
Figure 7: X-Gal staining
(A) X-Gal staining of E16.5 Selm-/- (a, b) and wild type mice (c, d). Wild type embryos were simultaneously stained with Alizarin red and Alcian blue to visualize bone and cartilage, respectively (e, f). Calvarial bones are marked by arrows. Scale bar, 2 mm.
(B) X-Gal staining of P4 wild type (+/+) and Selm-/- mice (-/-). Staining of the head (b), hindlimb (c), and ribs (d) of Selm-/- mice is shown in detail. Scale bar, 5 mm.
[*] Corresponding Author:
Prof. Dr. Andreas Winterpacht, Institute of Human Genetics, University Hospital Erlangen, Friedrich-Alexander-University Erlangen-Nürnberg, Schwabachanlage 10, D-91054 Erlangen, Germany; Tel. ++49-9131-852-2019; FAX: ++49-9131-852-3232, eMail: Andreas.Winterpacht@uk-erlangen.de